3. Data Format and Readers¶
3.1. CSV and TSV formats¶
The main formats used in GDSCTools are CSV-based but TSV-based formatted files may be accepted although not encouraged. The data files may be compressed using bz2, xz, gzip of zip methods. See Pandas documentation to know the exact list of authorised compression method. Note that files saved using GDSCTools will be saved in CSV format only.
GDSCTools provides tools to read different kind of structured CSV files. For
instance in the ANOVA analysis, these 3 types of CSV-input files defined in readers module) are used:
There are all based on the same Reader class. Not a
number values are encoded with the string NA or NaN. Besides, empty
strings or fields made of spaces or tabs are also replaced by spaces.
Quote characters are also removed.
Warning
Some readers will use the name of the extensions to infer the separator so it is important that the extension reflects the content of the file. A compressed file named as e.g. ic50.csv.gz will therefore be interpreted as a CSV file.
Note
With CSV files, if a column’s name is ambiguous in the header (i.e, contains already a comma), then it should be enclosed within double quotes to avoid ambiguities (e.g A,B should be “A,B”). See the Drug Decode section for an example.
3.2. IC50¶
The specific format for the IC50 data is CSV file where the header must contain a column named COSMIC_ID. Other column should correspond to a given drug identifier (an integer). The order of the columns does not matter. So, each row contains the IC50s for a given COSMIC identifier.
Warning
the data read in the CSV file is not transformed. So, IC50 data should be logged data.
Note
The IC50 matrix can be populated with other data (e.g., AUCs).
Although the name in the header does not matter note that if one column’s name starts with Drug then all columns that do not start with Drug are ignored (except the special column COSMIC_ID). This feature was implemented to account for old data files that stores all Drug identifiers and also all genomic features within a single file.
Here is an example of IC50 input:
COSMIC_ID, 1, 20, 40
111111, 0.5, 0.8, 10
222222, 1, 2, 10
The following old-style IC50 would be equivalent since (i) the last column is dropped (does not start with Drug) and (ii) the other column’s names are transformed into integer keeping only the middle part:
COSMIC_ID, Drug_1_IC50, Drug_20_IC50, Drug_40_IC50, other
111111, 0.5, 0.8, 10, 10
222222, 1, 2, 10, 20
If you save that example in a file (or download it _static/ic50_tiny.csv), you can read it with the
IC50 class as follows:
>>> from gdsctools import IC50
>>> r = IC50('_static/ic50_tiny.csv')
>>> r.drugIds
[1, 20, 40]
Note
the columns’ names should be identifiers (not drug names). There are two main reasons. The first one is that it allows us to keep anonymous all drug names and targets. The second reason is that many characteristics such as plate number and drug concentration may be associated with a drug identifier. This should be stored in a different table rather than in the name. It can then be handled and interpreted using the DrugDecode file (see below).
Note
column without a name are ignored.
See also
developers should look at the references for more
functionalities of the IC50
class (e.g., filter by tissues, removing drugs, visualisation of IC50s).
3.3. Genomic Features¶
The ANOVA analysis computes the associations between the IC50 and genomic features. This is the second input data set required for instance in the ANOVA analysis. Be aware that in the ANOVA analysis, the intersection between the IC50 and GenomicFeatures is made on the COSMIC_ID: cell lines not found in both CSV files will be dropped.
In addition to the COSMIC identifiers, the genomic feature file should contain the following columns:
- TISSUE_FACTOR
- MSI_FACTOR
- MEDIA_FACTOR
If not provided, the tissue, MSI and MEDIA factors will not be taken into account in the regression analysis. If the TCGA tissue is not provided, it is created and set to unidentified.
Note
Changed in version 0.9.11: A column called ‘Sample Name’ was interpreted if found. This is not the case anymore. It is actually removed now.
All remaining columns are assumed to be genomic features.
Warning
In the current version, all columns starting with Drug_ are removed without warning.
Here is a simple example:
COSMIC_ID, TISSUE_FACTOR, MSI_FACTOR, BRAF_mut, gain_cna
111111, lung_NSCLC, 1, 1, 0
222222, prostate, 1, 0, 1
It can be saved and read as follows with the GenomicFeatures
>>> from gdsctools import GenomicFeatures
>>> gf = GenomicFeatures('_static/gf_tiny.csv')
>>> gf
GenomicFeatures <Nc=2, Nf=2, Nt=2>
In GDSCTools, we provide a zipped Genomic Features file. It contains about 1000 cell lines and 47 genomic features (gene mutations). A more complex file tagged v17 is also provided with about 600 features v17 genomic feature.
Note that you may create instance of GenomicFeatures without input but a default data set is loaded (the subset aforementionned):
>>> from gdsctools import GenomicFeatures
>>> gf = GenomicFeatures()
>>> print(gf)
Genomic features distribution
Number of unique tissues 27
Here are the first 10 tissues: myeloma, nervous_system, soft_tissue, bone, lung_NSCLC, skin, Bladder, cervix, lung_SCLC, lung
MSI column: yes
MEDIA column: no
There are 47 unique features distributed as
- Mutation: 47
- CNA (gain): 0
- CNA (loss): 0
3.4. Combine IC50 and Genomic Features¶
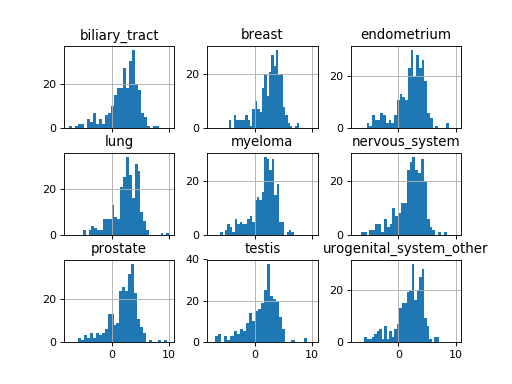
Here is an example on how to plot histograms of IC50s grouped by tissues. For convenience, we keep only 9 tissues.
from gdsctools import *
from numpy import mean
ic50 = IC50(ic50_v17)
gf = GenomicFeatures(gf_v17)
# select tissue column in same order as those stored in IC50 dataframe
tissues = gf.df.loc[ic50.df.index]['TISSUE_FACTOR']
ic50.df['tissue'] = tissues
# keep only 9 tissues
tokeep = list(set(tissues))[0:9]
ic50.df = ic50.df.query("tissue in @tokeep")
# Group by tissues
tt = ic50.df.groupby("tissue").aggregate(mean).transpose()
#plot histogram of IC50 group by tissues
tt.hist(bins=30, sharex=True)
(Source code, png, hires.png, pdf)
3.5. Drug Decode¶
DrugDecode files are not required to perform the analysis. You may skip that section.
Drugs used in GDSCTools analysis may be public or not. In order to guarantee that drugs are kept anonymised (if not public), we enforce the CSV files that contains the IC50s to used drug identifiers instead of drug names.
When creating reports, the Data Packages producer or owner or the drugs may want to decode the drug identifier. The information to perform that task is provided within the DrugDecode CSV file.
The DrugDecode class reads a CSV file that contains information about a drug and its target(s). It must contain 3 columns named as
follows:
DRUG_ID, DRUG_NAME, DRUG_TARGET
999, Erlotinib, EGFR
1039, SL 0101-1, "RSK, AURKB, PIM3"
Note the usage of quotes in the last row/last columns to avoid conflicts with the CSV format itself.
These columns will be used if provided:
- WEBRELEASE
- OWNED_BY
In addition, these columns may be populated for later use:
- CHEMSPIDER_ID
- PUBCHEM_ID
- CHEMBL_ID
An example can be read as follows:
>>> from gdsctools import DrugDecode, datasets
>>> drug_filename = datasets.testing.drug_test_csv.location
>>> dd = DrugDecode(drug_filename)
>>> dd.get_name(1047)
'Nutlin-3a'
>>> dd.df.loc[999]
CHEMBL_ID NaN
CHEMSPIDER_ID NaN
DRUG_NAME Erlotinib
DRUG_TARGET EGFR
OWNED_BY NaN
PUBCHEM_ID NaN
WEBRELEASE NaN
Name: 999, dtype: object
DrugDecode files are not required for the analysis but are used by
gdsctools.anova_report.ANOVAReport to fill the HTML reports.
You can also run the analysis and set the drug names and target later on as
follows using the drug_annotations method:
from gdsctools import *
an = ANOVA(ic50_test)
an.anova_all()
results = an.anova_all()
dd = DrugDecode("v19_drug_decode.csv")
newdf = dd.drug_annotations(results.df)